Backbone-to-Sequence Inverse Folding
Studies how geometric structure encoding and sequence decoding recover amino-acid sequences from protein backbones.
Description
Task: Protein Inverse Folding — Structure Encoder Design
Research Question
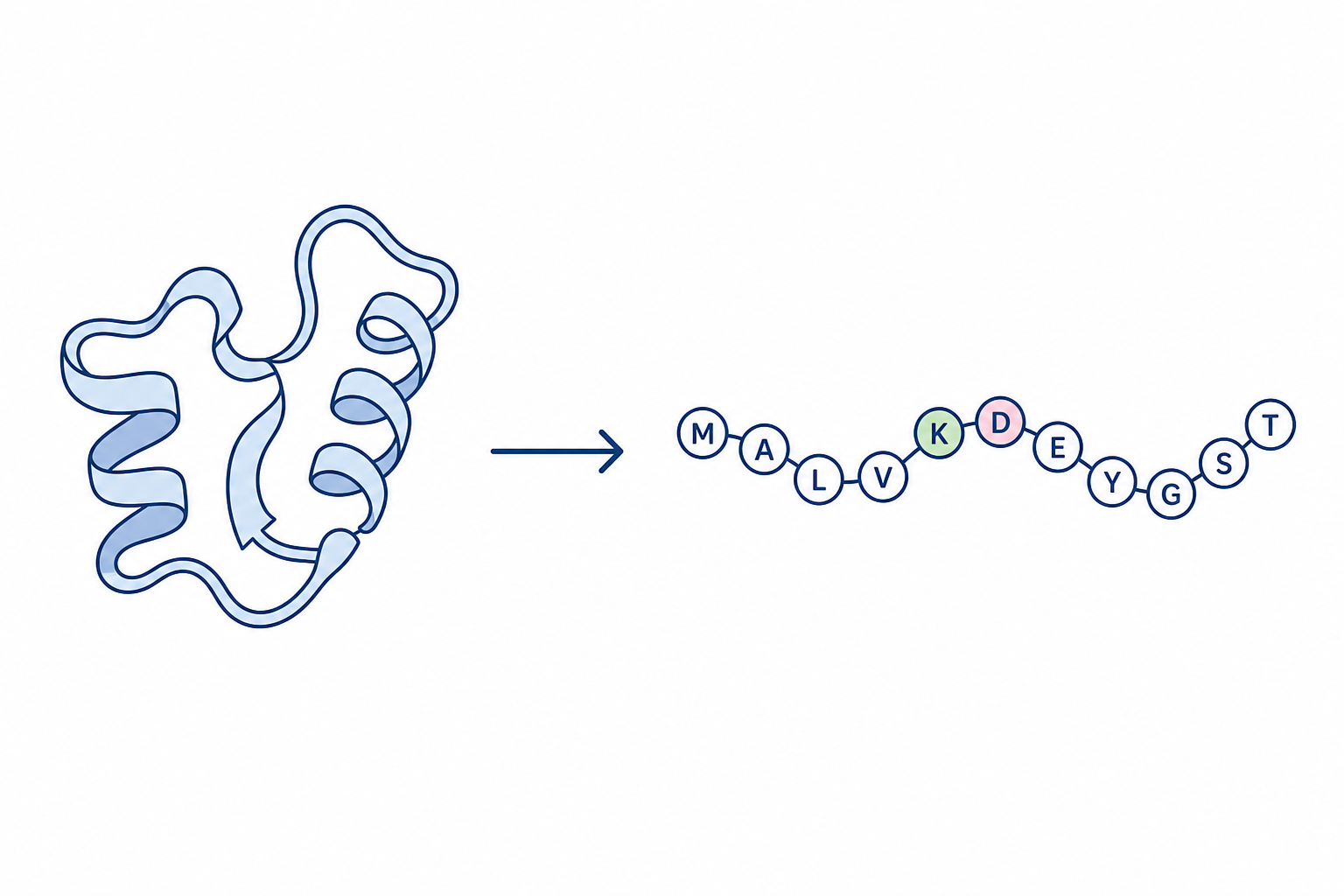
Design a novel GNN-based structure encoder for protein inverse folding: given backbone atom coordinates (N, CA, C, O), predict the amino acid sequence that would fold into that structure.
Background
Protein inverse folding (also called computational protein design or fixed-backbone design) is a central problem in structural biology. Given a protein backbone structure, the goal is to predict the amino acid sequence most likely to fold into that structure. This is the inverse of the protein folding problem (predicting structure from sequence).
The key challenge is encoding the 3D protein backbone graph into rich per-residue embeddings that capture local geometry, long-range interactions, and structural motifs. Existing approaches differ primarily in how they encode the protein structure:
- GVP (Geometric Vector Perceptron; Jing et al., "Learning from Protein Structure with Geometric Vector Perceptrons", ICLR 2021; arXiv:2009.01411). SE(3)-equivariant message passing with both scalar and vector node/edge features. Code: https://github.com/drorlab/gvp.
- ProteinMPNN (Dauparas et al., "Robust deep learning–based protein sequence design using ProteinMPNN", Science 2022, 378(6615):49–56; bioRxiv 2022.06.03.494563). Message-passing encoder with edge updates, followed by an autoregressive decoder with masking. Code: https://github.com/dauparas/ProteinMPNN.
- PiFold (Gao et al., "PiFold: Toward Effective and Efficient Protein Inverse Folding", ICLR 2023; arXiv:2209.12643). PiGNN encoder with learnable virtual atoms, multi-scale distance features, and dihedral features, plus a non-autoregressive one-shot decoder. Code: https://github.com/A4Bio/PiFold.
The structure encoder is the critical component: all methods share the same input format (backbone coordinates) and output format (amino acid log-probabilities), but differ in how they transform structure into sequence-informative representations.
What to Implement
Modify the editable section of custom_invfold.py. You must implement:
- StructureEncoder: A GNN module that takes backbone coordinates
X(B, L, 4, 3) and mask (B, L), and produces per-residue embeddingsh_V(B, L, hidden_dim). - InverseFoldingModel: Wraps the encoder with a decoder head that outputs amino acid log-probabilities (B, L, 20).
Interface
class StructureEncoder(nn.Module):
def __init__(self, hidden_dim=128, ...):
...
def forward(self, X, mask):
"""
X: (B, L, 4, 3) backbone coordinates [N, CA, C, O]
mask: (B, L) binary mask (1 for valid residues, 0 for padding)
Returns: h_V (B, L, hidden_dim) per-residue embeddings
"""
...
class InverseFoldingModel(nn.Module):
def __init__(self, hidden_dim=128, ...):
...
def forward(self, X, mask):
"""
Returns: log_probs (B, L, 20) amino acid log-probabilities
"""
...Helper functions available in the FIXED section above the editable region:
_rbf(D, ...): Radial basis function encoding of distances._dihedrals(X): Backbone dihedral angles (phi, psi, omega) as sin/cos features._orientations(X): Local coordinate frame (forward + binormal vectors).knn_graph(X_ca, mask, k): Build k-nearest neighbor graph from CA coordinates.
Fixed Pipeline
Datasets, train/validation/test splits, the training loop, padding/masking, optimizer schedule, loss (per-residue cross-entropy), and evaluation harness are all supplied by the scaffold and not part of the contribution.
Evaluation
The model is evaluated on three benchmarks:
- CATH 4.2: Standard protein design benchmark (single-chain, ~18k train / 608 test).
- CATH 4.3: Updated CATH with more diverse structures (~21k train / 1120 test).
- TS50: 50 de novo designed proteins for out-of-distribution generalization (trained on CATH 4.2).
Primary metric: Recovery (fraction of correctly predicted amino acids, higher is better). Secondary metric: Perplexity (exponential of per-residue cross-entropy loss, lower is better).
Code
1"""2Protein Inverse Folding — Self-contained template.3Given backbone structure (N, CA, C, O coordinates), predict amino acid sequence.45Structure:6Lines 1-75: FIXED — Imports, constants, data loading, featurization7Lines 76-230: EDITABLE — StructureEncoder + decoder (starter: simple MPNN)8Lines 231+: FIXED — Training loop, evaluation, metrics9"""10import os11import sys12import json13import math14import time15import argparse
Method Summary
ProteinMPNN + RLFE + GeoGate
Augments ProteinMPNN edges with relative local-frame rotations and gates messages by frame-conditioned sigmoids.